Nanolive is proud to be part of a research collaboration with leading virologists from the group of Olivier Schwartz at the Institut Pasteur in Paris. The latest research from Nanolive’s Deep Quantitative Biology department and the Institut Pasteur uses cutting-edge AI-based computer vision to give new insights into organelle network disruption during coronavirus infection. The group have published their findings in an open access pre-print available on bioRxiv: “Dynamic label-free analysis of SARS-CoV-2 infection reveals virus-induced subcellular remodeling”.
The Head of AI for Quantitative Biology at Nanolive, Mathieu Fréchin shared his enthusiasm for the opportunity to collaborate with the Institut Pasteur: “Working with Nell, Olivier, Tim and all the other members of Pasteur was the greatest chance for the Deep Quantitative department to develop an original quantitative biology approach to understand SARS-CoV-2 infection. We created bridges that we hope will nurture novel AI-fuelled research.”
Label-free analysis of organelles
Building on the expertise used to develop Nanolive’s existing digital assays for cellular image analysis, members of Nanolive’s Deep Quantitative Biology team created new automated methods for detecting and quantifying cell nuclei, nucleoli and mitochondria in label-free holotomographic images (see video and images below). These digital segmentations of organelles could then be used to quantify organelle numbers, their changing morphology, interactions, mass, and other features.
Timelapse video showing U2OS-ACE2 cells infected with SARS-CoV-2 Omicron BA.1. The version on the left includes the digital segmentation overlay of organelle masks; nuclei (cyan), nucleoli (pink), mitochondria (red) and lipid droplets (green). The version on the right shows the refractive index signal only. Supplementary video 16 from Saunders et. al, bioRxiv 2023.11.16.567378.
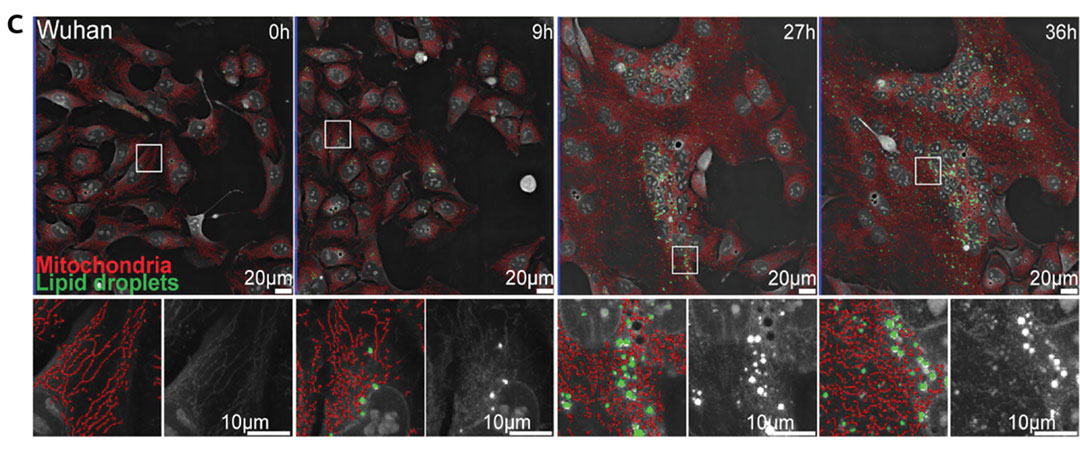
Images showing the digital segmentation of mitochondria (red) and lipid droplets (green) demonstrate the accuracy of the analysis method. Digital segmentations are shown overlaid on holotomographic images where objects with a higher refractive index appear brighter. Extract from Figure 3C, Saunders et. al, bioRxiv 2023.11.16.567378.
Subcellular remodelling revealed
The automated detection and segmentation of organelles allowed researchers at the Institut Pasteur to detect and analyse how networks of organelles interacted and changed during coronavirus infection. Cells were imaged for 36 hours, after exposure to different SARS-CoV-2 strains. The results indicate a wide range of changes in cells infected with SARS-CoV-2, including cell-cell fusion, fragmentation of mitochondrial networks, perinuclear localization of lipid droplets, and increased nucleoli size.
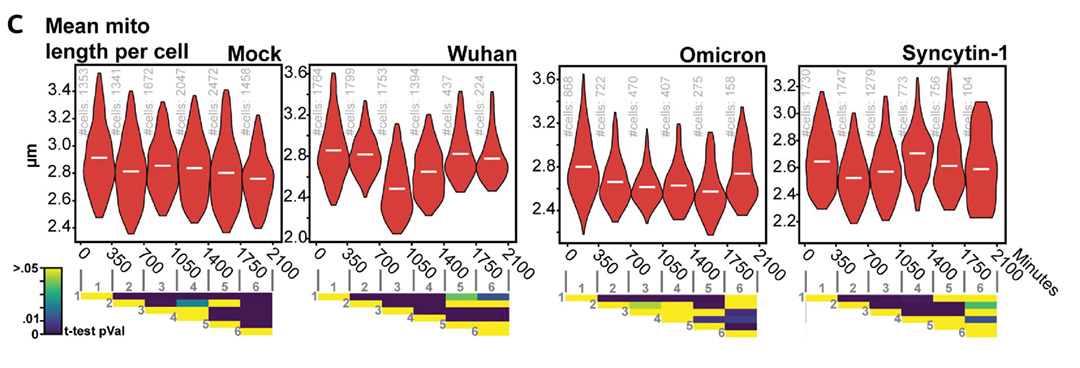
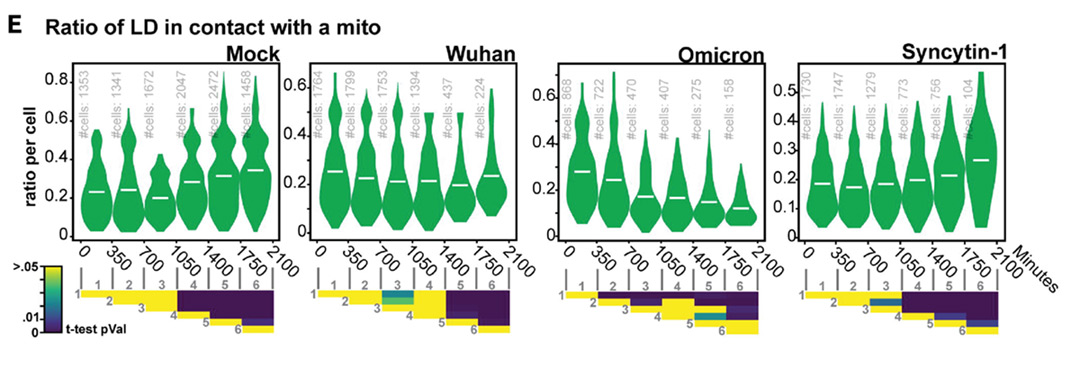
Two of the measurements produced were the mean mitochondrion length per cell (top), and the ratio of lipid droplets (LD) in contact with a mitochondrion per cell (bottom), shown for different conditions. Extract from Figure 4C and 4E, Saunders et. al, bioRxiv
The next generation of phenotypic image analysis
This innovative work by the Nanolive team is at the forefront of research and as such, is not part of the automated digital analysis assays currently offered on the Nanolive platform.
In the future, we hope that by visualizing organelles and their dynamics label-free, AI-analysis for quantitative holotomography will ensure that results are not biased by the effects of chemical or fluorescent labels, which particularly affect mitochondrial networks, and sensitive cell types. Additionally, by studying responses across many organelles simultaneously as demonstrated in this paper, the chance of missing key effects can be reduced. In combination with Nanolive’s high resolution, label-free images, this approach could be a promising solution for quantitative phenotypic screening at cellular and organelle levels, and has the potential to give unique insights into dynamic cellular responses to drugs and disease at a level of detail that remains out of reach for other imaging technologies.
Read the full paper on bioRxiv.
Nanolive’s Smart Lipid Droplet AssayLIVE was used to segment and quantify lipid droplets in this paper. Nucleus, nucleoli, and mitochondrial segmentation are not currently available in the Nanolive digital assay products.
To find out more about Nanolive’s holotomogaphic imaging systems, click here.
Read our latest news
Cytotoxic Drug Development Application Note
Discover how Nanolive’s LIVE Cytotoxicity Assay transforms cytotoxic drug development through high-resolution, label-free quantification of cell health and death. Our application note explores how this advanced technology enables real-time monitoring of cell death...
Investigative Toxicology Application Note
Our groundbreaking approach offers a label-free, high-content imaging solution that transforms the way cellular health, death, and phenotypic responses are monitored and quantified. Unlike traditional cytotoxicity assays, Nanolive’s technology bypasses the limitations...
Phenotypic Cell Health and Stress Application Note
Discover the advanced capabilities of Nanolive’s LIVE Cytotoxicity Assay in an application note. This document presents a detailed exploration of how our innovative, label-free technology enables researchers to monitor phenotypic changes and detect cell stress...



