We are delighted to announce that a new publication featuring Nanolive imaging has been published in BMC Molecular and Cell Biology. The study, a huge international collaboration spearheaded by Dr. Michael Cahill from the Charles Sturt University (NSW, Australia) and published by lead authors Bashar Thejer and Partho Adhikary, used genetically-modified human pancreatic cancer cells (MIA-PaCa-2) to investigate the effect phosphorylation status of the Progesterone Receptor Membrane Component 1 (PGRMC1) has on cell metabolism and morphology.
Read other scientific publications using Nanolive’s imaging here.
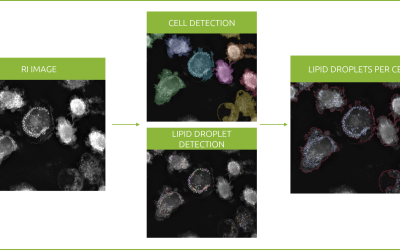
PGRMC1 is a heme-binding protein that plays an important role in membrane trafficking, cell cycling and lipid transport. Significant upregulation of PGRMC1 has been observed in many cancers. Mutating the phosphorylation status of PGRMC1, resulted in epigenetic changes in the genome of MIA PaCa-2 cells, modifying cell differentiation and thus, morphology. This was accompanied by dramatic changes in mitochondria.
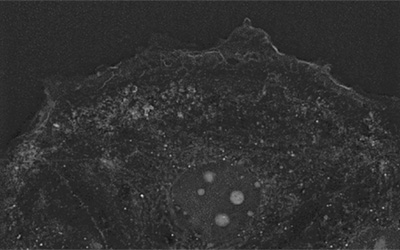
Dr. Cahill was kind enough to send us the footage that was taken using Nanolive imaging and discuss what we can see. In the wildtype (WT) population, we see a characteristic MIA PaCa-2 cell population, comprised of “fibroblast-shaped” cells, with a minority of cells displaying a more rounded morphology. All cells show the membrane blebbing characteristic of MIA PaCa-2 cells. It is possible that this fascinating behaviour is related to exosome secretion.
If we compare this footage to those in the S57A/Y180F/S181A triple mutant (TM), the differences are striking. TM cells are much rounder, and their mitochondria are much more fragmented than the WT cells. Proteomic pathway enrichment analysis confirmed that mitochondrial function was impaired in S57A/S181A double mutant (DM) and TM cells. This is probably caused by changes in the actin cytoskeleton, which is also strongly affected by the phosphorylation status of PGRMC1.
In the TM video, some of the cells appear to be tri-lobulated cells (i.e. have three nuclei). We asked Dr. Cahill whether this could be a treatment effect or whether this was normal for MIA PaCa-2 cells? “MIA PaCa cells are a bit of a disaster; the genome is so unstable that cytologists are often unable to identify the chromosomes from two different cells in the same culture. During mitosis, cells get to cytokinesis but they sometimes never complete it. Instead of separating, they rejoin and then you have one cell with triple the genome.” It is not known how frequently this type of event occurs, but Dr. Cahill speculates that it could be influenced by PGRMC1. “Others have shown that the protein itself is located on the spindle in mitosis and that it regulates whether cells enter the cell cycle or the resting phase of the cell cycle (G0), so it is likely to have some kind of overarching role there but this has not been very well characterized to date”.
Investigating such hypotheses with Nanolive imaging is feasible and could be an extremely interesting follow up study. Nanolive thanks Dr. Cahill for taking the time to speak to us and congratulate all the authors on an excellent and important study.
Access our latest Oncology and Immuno-Oncology application note here.
Read our latest news
Revolutionizing lipid droplet analysis: insights from Nanolive’s Smart Lipid Droplet Assay Application Note
Introducing the Smart Lipid Droplet Assay: A breakthrough in label-free lipid droplet analysis Discover the power of Nanolive's Smart Lipid Droplet Assay (SLDA), the first smart digital assay to provide a push-button solution for analyzing lipid droplet dynamics,...
Food additives and gut health: new research from the University of Sydney
The team of Professor Wojciech Chrzanowski in the Sydney Pharmacy School at the University of Sydney have published their findings on the toxic effect of titanium nanoparticles found in food. The paper “Impact of nano-titanium dioxide extracted from food products on...
2023 scientific publications roundup
2023 has been a record year for clients using the Nanolive system in their scientific publications. The number of peer-reviewed publications has continued to increase, and there has been a real growth in groups publishing pre-prints to give a preview of their work....
Nanolive microscopes

3D CELL EXPLORER
Budget-friendly, easy-to-use, compact solution for high quality non-invasive 4D live cell imaging

3D CELL EXPLORER-fluo
Multimodal Complete Solution: combine high quality non-invasive 4D live cell imaging with fluorescence

CX-A
Automated live cell imaging: a unique walk-away solution for long-term live cell imaging of single cells and cell populations